SMAD4 is detected as a mutational cancer driver
SMAD4 reports
Gene details
| SMAD4 |
|---|
|
Ensembl ID
ENSG00000141646
Transcript ID ENST00000342988 Protein ID ENSP00000341551 |
|
Cancer types where is driver
13
Cohorts where is driver 35 Mutated samples 552 Mutations 872 |
| Mode of action Ambiguous |
| Known driver True |
Method signals per Cancer Type
| Cancer type | Methods | Samples | Samples (%) |
|---|
ClustL
HotMAPS
smRegions
Clustered Mutations
CBaSE dNdScv Recurrent Mutations
FML Functional Mutations
MutPanning Tri-nucleotide specific bias
combination Combination
CBaSE dNdScv Recurrent Mutations
FML Functional Mutations
MutPanning Tri-nucleotide specific bias
combination Combination
In-silico saturation mutagenesis
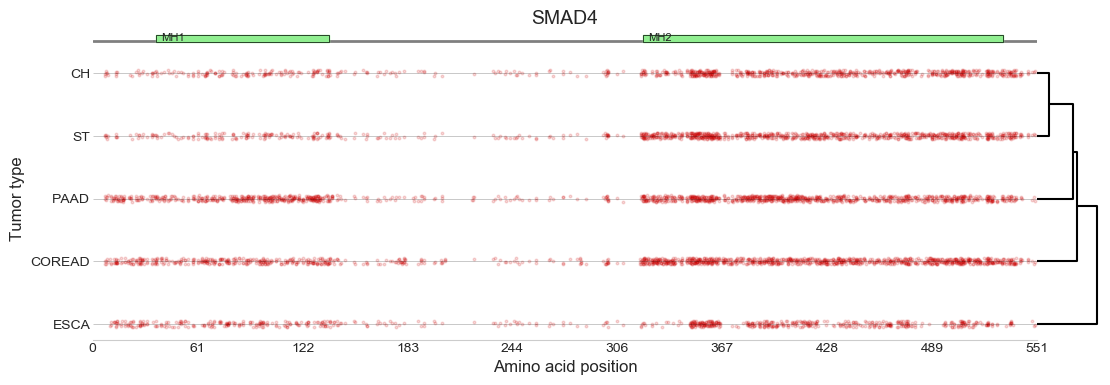
In silico saturation mutagenesis profiles across all tumor types
where a specific model is available.
Driver mutations (displayed as red dots) are mapped to
their positions in the canonical transcript.
The dendrogram on the right represents a hierarchical
clustering of the profiles based on site-by-site
concordance via Matthews Correlation similarity.
The track on the top displays the Pfam domains mapping
to the canonical transcript.
See the FAQs for more details
about the in silico saturation mutagenesis computed
with boostDM.
| Model | Mutations | Driver mutations | Driver mutations (%) |
|---|---|---|---|
| COREAD (Colorectal Adenocarcinoma) | 4959 | 1319 | 26.60 |
| ESCA (Esophageal Adenocarcinoma) | 4959 | 706 | 14.24 |
| PAAD (Pancreatic Adenocarcinoma) | 4959 | 1357 | 27.36 |
Observed mutations in tumors
The mutations needle plot shows the distribution of the observed
mutations along the protein sequence.
| Mutation (GRCh38) | Protein Position | Samples | Consequence |
|---|